Introduction on Molecular Beacons
Molecular beacons are single-stranded oligonucleotide hybridization probes that form a stem-and-loop structure. The loop contains a probe sequence that is complementary to a target sequence, and the stem is formed by the annealing of complementary arm sequences that are located on either side of the probe sequence. A fluorophore is covalently linked to the end of one arm and a quencher is covalently linked to the end of the other arm. Molecular beacons do not fluoresce when they are free in solution. However, when they hybridize to a nucleic acid strand containing a target sequence they undergo a conformational change that enables them to fluoresce brightly.
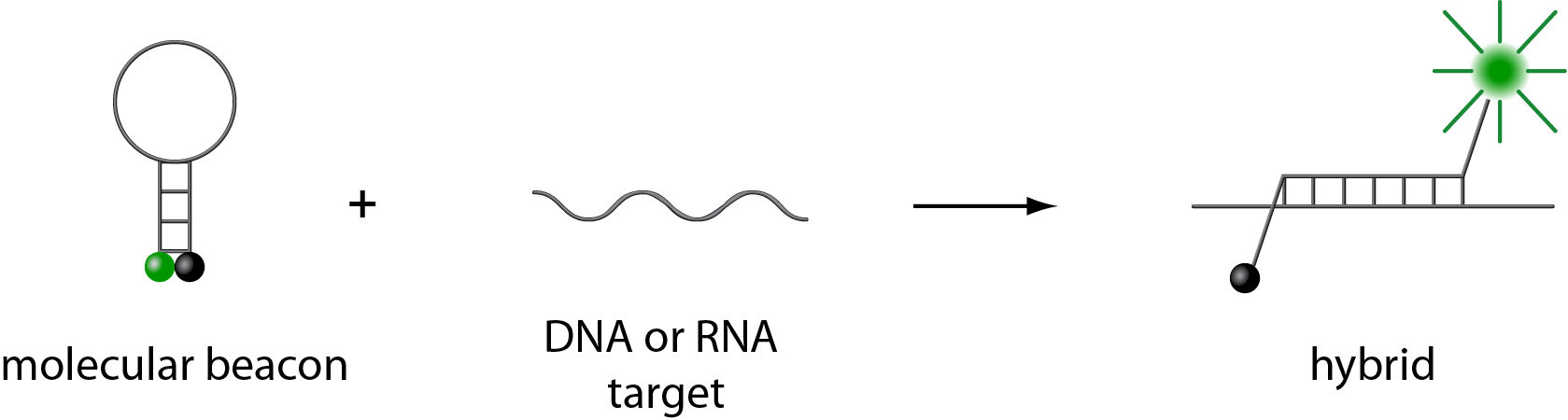
In the absence of targets, the probe is dark, because the stem places the fluorophore so close to the nonfluorescent quencher that they transiently share electrons, eliminating the ability of the fluorophore to fluoresce. When the probe encounters a target molecule, it forms a probe-target hybrid that is longer and more stable than the stem hybrid. The rigidity and length of the probe-target hybrid precludes the simultaneous existence of the stem hybrid. Consequently, the molecular beacon undergoes a spontaneous conformational reorganization that forces the stem hybrid to dissociate and the fluorophore and the quencher to move away from each other, restoring fluorescence.
Molecular beacons can be used as amplicon detector probes in diagnostic assays. Because nonhybridized molecular beacons are dark, it is not necessary to isolate the probe-target hybrids to determine the number of amplicons synthesized during an assay. Molecular beacons are added to the assay mixture before carrying out gene amplification and fluorescence is measured in real time. The assay tube remains sealed. Consequently, the amplicons cannot escape to contaminate untested samples. Furthermore, the use of molecular beacons provides an additional level of specificity. Because it is very unlikely that false amplicons or primer-dimers possess target sequences for the molecular beacons, the generation of fluorescence is exclusively due to the synthesis of the intended amplicons.
Molecular beacons can be synthesized that possess differently colored fluorophores, enabling assays to be carried out that simultaneously detect different targets in the same reaction. For example, multiplex assays can contain a number of different primer sets, each set enabling the amplification of a unique gene sequence from a different pathogenic agent, and a corresponding number of molecular beacons can be present, each containing a probe sequence specific for one of the amplicons, and each labeled with a fluorophore of a different color. The color of the resulting fluorescence, if any, identifies the pathogenic agent in the sample, and the number of amplification cycles required to generate detectable fluorescence provides a quantitative measure of the number of target organisms present. If more than one type of pathogen is present in the sample, the fluorescent colors that occur identify which are present. Moreover, due to the inherent design of gene amplification assays, the use of molecular beacons enables the abundance of a rare pathogen to be determined in the presence of a much more abundant pathogen.
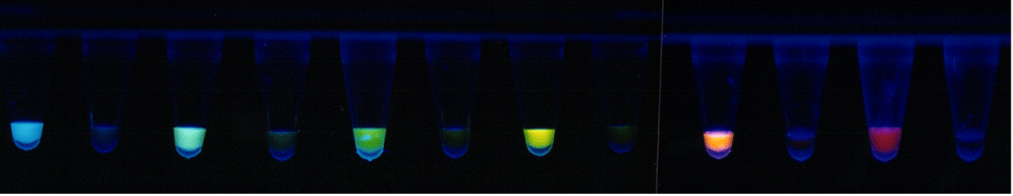
Molecular beacons are extraordinarily specific. They easily discriminate target sequences that differ from one another by a single nucleotide substitution. The reason that molecular beacons are so "finicky" is that they can exist in two different stable physical states. In one state, the molecular beacons are hybridized to their targets, and energy is stored in the probe-target helix. In the second state, the molecular beacons are free in solution, and energy is stored in their stem helix. Molecular beacons are designed so that their probe sequence is just long enough for a perfectly complementary probe-target hybrid to be more stable than the stem hybrid. Consequently, the molecular beacons spontaneously form fluorescent probe-target hybrids. However, if as little as a single nucleotide in the target is not complementary to the probe sequence of the molecular beacon, the probe-target helix would be less stable. In this situation, the stem helix of the molecular beacon is more stable than the mismatched probe-target helix, and the molecular beacons remain unhybridized. Thus, molecular beacons can be thought of as "molecular switches" that are on their targets and brightly fluorescent when the targets are perfectly complementary to the probe, but remain off the targets and dark if the targets contain a mutation.
Molecular beacons are thus ideal probes for use in diagnostic assays designed for genetic screening, SNP detection, and pharmacogenetic applications. For example, to determine the genotype of an individual at a particular locus, the genetic region of interest is amplified in the presence of two different molecular beacons, one perfectly complementary to the wild-type allele and labeled with a fluorophore of a particular color, and the other perfectly complementary to the mutant allele and labeled with a differently colored fluorophore. If the assay results in the generation of only the first fluorescent color, then the individual is homozygous wild type at that locus. If the assay results in the generation of only the other fluorescent color, then the individual is homozygous mutant. And finally, if both fluorescent colors are produced, then the individual is heterozygous.
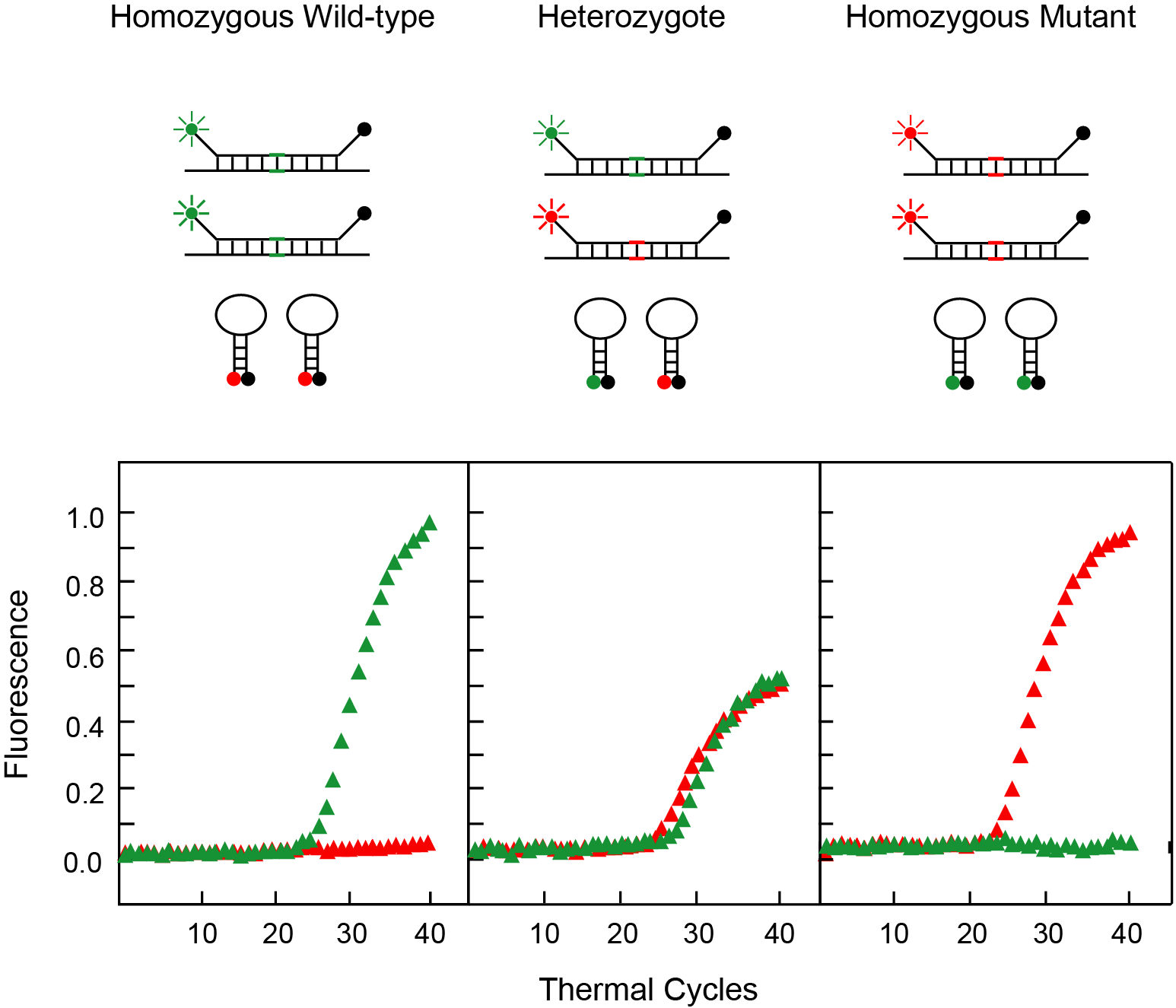
In summary, molecular beacons have three key properties that enable the design of new and powerful diagnostic assays: 1) they only fluoresce when bound to their targets, 2) they can be labeled with a fluorophore of any desired color, and 3) they are so specific that they easily discriminate single-nucleotide polymorphisms. Now that a number of new and versatile spectrofluorometric thermal cyclers are available to clinical diagnostic and research laboratories, assays that simultaneously utilize as many as seven differently colored molecular beacons can be designed. This enables cost-efficient multiplex assays to be developed that identify which member of a panel of potential infectious agents is present in a clinical sample. Utilizing molecular beacons, assays can be developed that not only identify a causative infectious agent, they can simultaneously determine which antibiotics will be effective and which will be ineffective against the particular strain that is present. A panel of different genetic mutations responsible for the same disease can be identified in a single assay. And a single assay containing molecular beacons can screen an entire panel of SNPs that determine whether a particular drug will be effective for a particular individual. Thus, molecular beacons provide a new tool for increasing the effectiveness and lowering the cost of clinical diagnostic assays.
Recent Publications from our group
Ma MT, Jiang Q, Chen CH, Badeti S, Wang X, Zeng C, Evans D, Bodnar B, Marras SAE, Tyagi S, Bharaj P, Yehia G, Romanienko P, Hu W, Liu SL, Shi L, and Liu D (2024) S309-CAR-NK cells bind the Omicron variants in vitro and reduce SARS-CoV-2 viral loads in humanized ACE2-NSG mice. Journal of Virology: e0003824. PMID: 38767356: PubMed Link
Banada PP, Green R, Streck D, Kurathi R, Reiss R, Banik S, Montalvan I, Jones R, Marras SAE, Chakravorty S, and Alland D (2023) An expanded RT-PCR melting temperature coding assay to rapidly identify all known SARS-CoV-2 variants and sub-variants of concern. Scientific Reports 13. 21927. PMID: 38081834: PubMed Link
Ebraham L, Xu C, Wang A, Hernandez C, Siclari N, Rajah D, Walter L, Marras SAE, Tyagi S, Fine DH, Daep CA, and Chang TL (2023) Oral Epithelial cells expressing low or undetectable levels of human angiotensin-converting enzyme 2 are susceptible to SARS-CoV-2 virus infection in vitro. Pathogens 12. PMID: 37375533: PubMed Link